A Molecular Switch Abrogates Glycoprotein 100 (gp100) T-cell Receptor (TCR) Targeting of a Human Melanoma Antigen.
Bianchi, V., Bulek, A., Fuller, A., Lloyd, A., Attaf, M., Rizkallah, P.J., Dolton, G., Sewell, A.K., Cole, D.K.(2016) J Biological Chem 291: 8951-8959
- PubMed: 26917722
- DOI: https://doi.org/10.1074/jbc.M115.707414
- Primary Citation of Related Structures:
5EU3, 5EU4, 5EU5, 5EU6 - PubMed Abstract:
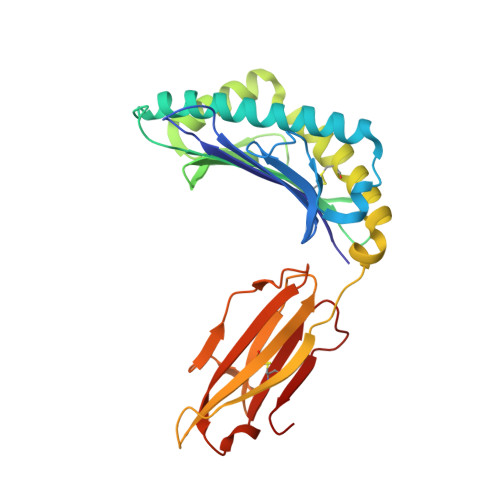
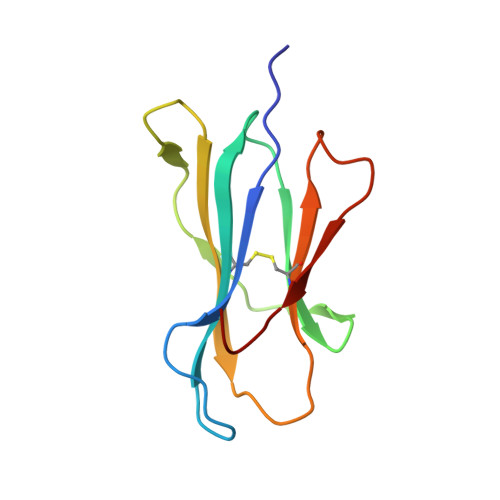
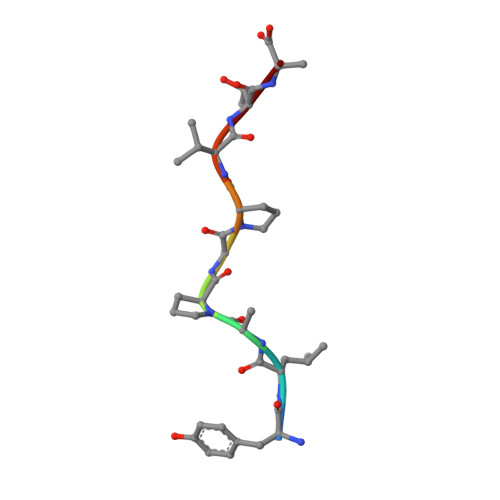
Human CD8(+) cytotoxic T lymphocytes can mediate tumor regression in melanoma through the specific recognition of HLA-restricted peptides. Because of the relatively weak affinity of most anti-cancer T-cell receptors (TCRs), there is growing emphasis on immunizing melanoma patients with altered peptide ligands in order to induce strong anti-tumor immunity capable of breaking tolerance toward these self-antigens. However, previous studies have shown that these immunogenic designer peptides are not always effective. The melanocyte differentiation protein, glycoprotein 100 (gp100), encodes a naturally processed epitope that is an attractive target for melanoma immunotherapies, in particular peptide-based vaccines. Previous studies have shown that substitutions at peptide residue Glu(3) have a broad negative impact on polyclonal T-cell responses. Here, we describe the first atomic structure of a natural cognate TCR in complex with this gp100 epitope and highlight the relatively high affinity of the interaction. Alanine scan mutagenesis performed across the gp100(280-288) peptide showed that Glu(3) was critically important for TCR binding. Unexpectedly, structural analysis demonstrated that the Glu(3) → Ala substitution resulted in a molecular switch that was transmitted to adjacent residues, abrogating TCR binding and T-cell recognition. These findings help to clarify the mechanism of T-cell recognition of gp100 during melanoma responses and could direct the development of altered peptides for vaccination.
- From the Division of Infection and Immunity and Systems Immunity Research Institute, Cardiff University School of Medicine, Heath Park, Cardiff CF14 4XN, United Kingdom.
Organizational Affiliation: