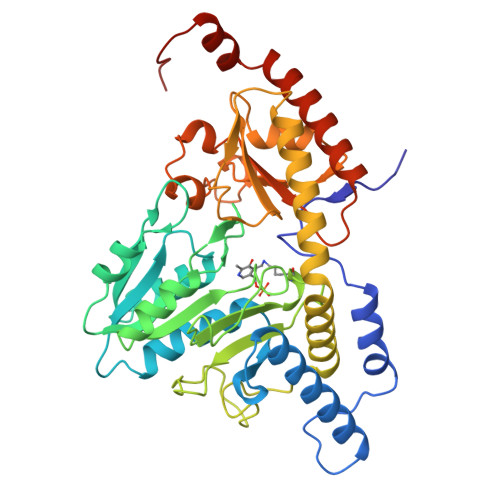
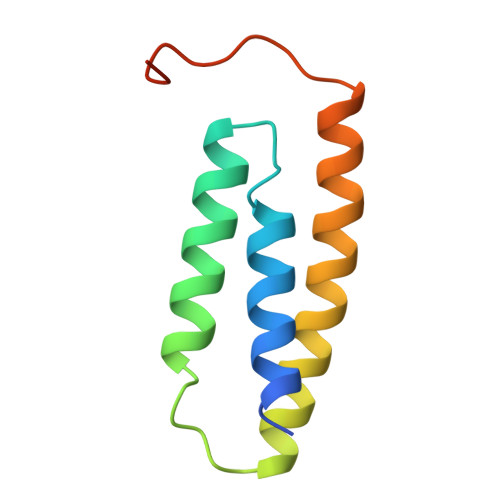
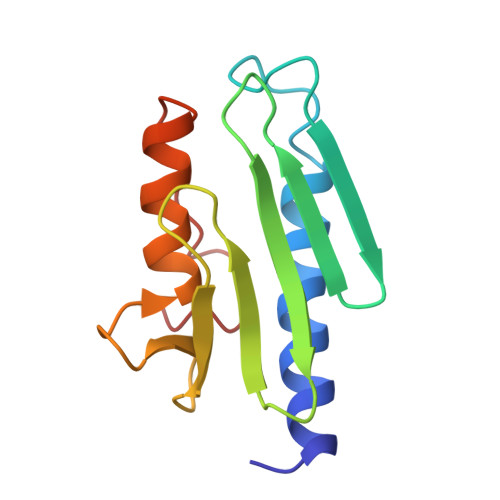
Architectural Features of Human Mitochondrial Cysteine Desulfurase Complexes from Crosslinking Mass Spectrometry and Small Angle X-ray Scattering
Cai, K., Frederick, R.O., Dashti, H., Markley, J.L.(2018) Structure 26: 1127-1136
- PubMed: 29983374
- DOI: https://doi.org/10.1016/j.str.2018.05.017
- Primary Citation of Related Structures:
8ZZF - PubMed Abstract:
Cysteine desulfurase plays a central role in mitochondrial iron-sulfur cluster biogenesis by generating sulfur through the conversion of L-cysteine to L-alanine and by serving as the platform for assembling other components of the biosynthetic machinery, including ISCU, frataxin, and ferredoxin. The human mitochondrial cysteine desulfurase complex consists of two copies each of NFS1, ISD11, and acyl carrier protein. We describe results from chemical crosslinking coupled with tandem mass spectrometry and small-angle X-ray scattering studies that are consistent with a closed NFS1 dimer rather than an open one for both the cysteine desulfurase-ISCU and cysteine desulfurase-ISCU-frataxin complexes. We present a structural model for the cysteine desulfurase-ISCU-frataxin complex derived from chemical crosslinking restraints in conjunction with the recent crystal structure of the cysteine desulfurase-ISCU-zinc complex and distance constraints from nuclear magnetic resonance.
- Biochemistry Department, University of Wisconsin-Madison, 433 Babcock Drive, Madison, WI 53706, USA.
Organizational Affiliation: