Biochemical, Structural, and Conformational Characterization of a Fungal Ethylene-Forming Enzyme.
Chatterjee, S., Rankin, J.A., Farrugia, M.A., J S Rifayee, S.B., Christov, C.Z., Hu, J., Hausinger, R.P.(2025) Biochemistry 64: 2054-2067
- PubMed: 40052306
- DOI: https://doi.org/10.1021/acs.biochem.5c00038
- Primary Citation of Related Structures:
9EIR, 9EIS - PubMed Abstract:
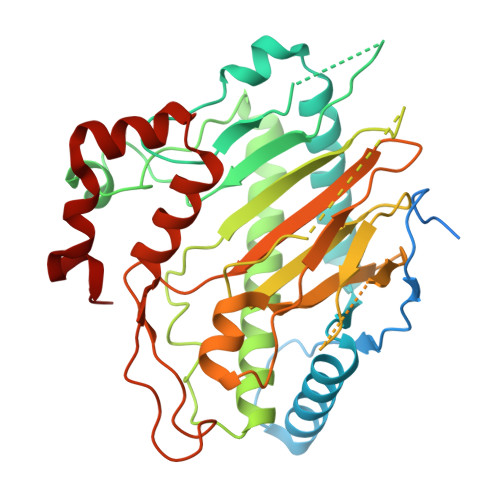
The ethylene-forming enzyme (EFE) from the fungus Penicillium digitatum strain Pd1 was heterologously produced in Escherichia coli and its properties were compared to the extensively characterized bacterial enzyme from Pseudomonas savastanoi strain PK2. Both enzymes catalyze four reactions: the conversion of 2-oxoglutarate (2OG) to ethylene and CO 2 , oxidative decarboxylation of 2OG coupled to l-arginine (l-Arg) hydroxylation, uncoupled oxidative decarboxylation of 2OG, and the production of 3-hydroxypropionate (3-HP) from 2OG. The strain Pd1 enzyme exhibited a greater ratio of ethylene production over l-Arg hydroxylation than the PK2 strain EFE. The uncoupled decarboxylation of 2OG and 3-HP production are minor reactions in both cases, but they occur to a greater extent using the fungal enzyme. Additional distinctions of the fungal versus bacterial enzyme are noted in the absorbance maxima and l-Arg dependence of their anaerobic electronic spectra. The structures of the Pd1 EFE apoprotein and the EFE·Mn(II)·2OG complex resembled the corresponding structures of the PK2 enzyme, but notable structural differences were observed in the computationally predicted Pd1 EFE·Fe(II)·2OG·l-Arg complex versus the PK2 EFE·Mn(II)·2OG·l-Arg crystal structure. These studies extend our biochemical understanding and represent the first structural and conformational characterization of a eukaryotic EFE.
- Department of Microbiology and Molecular Genetics, Michigan State University, East Lansing, Michigan 48824, United States.
Organizational Affiliation: