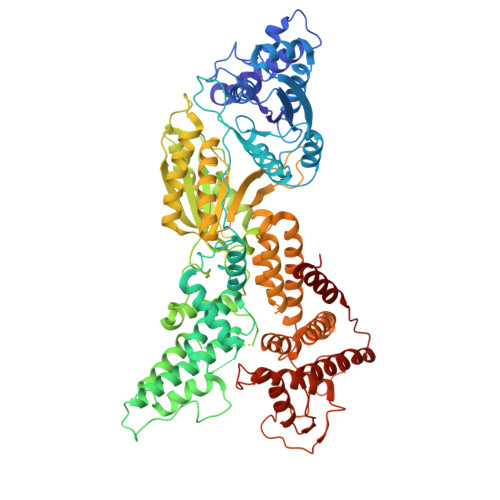
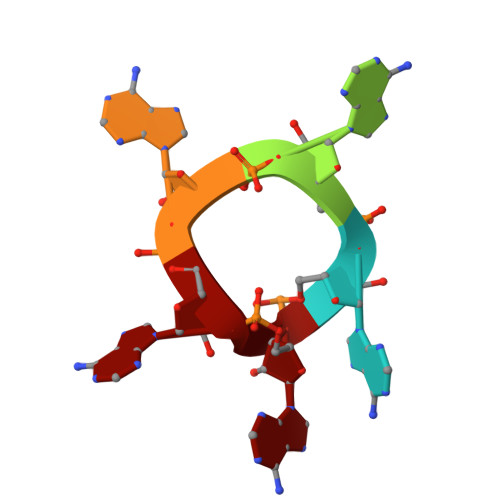
Mechanistic basis for selective Csm6-2 activation by cyclic penta-adenylate in a type III CRISPR-Cas system.
Shi, R., Yang, M., Liu, Y., Gao, H., Lin, Z.(2026) EMBO J
- PubMed: 41917262 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s44318-026-00767-3
- Primary Citation Related Structures:
9W3U, 9W3V, 9W3W - PubMed Abstract:
Type III CRISPR systems generate cyclic oligoadenylate (cOA, 3 to 6 AMPs) messengers upon detecting viral RNA, activating downstream effectors to defend against viral infection. Although cOA-activated effectors have been extensively characterized, the effectors specific to cA5-one of the most abundant cOA species produced during phage infection-have remained unexplored. Here, we report that the CRISPR ribonuclease Csm6 (Csm6-2) from Actinomyces procaprae selectively employs cA5 as its activator. Csm6-2 utilizes its HEPN domain, rather than the CARF domain, to mediate self-limiting cleavage of cOA activators. Cryo-EM structural analyses reveal that Csm6-2 functions as a homotetramer, and disruption of tetramer formation significantly reduces its ribonuclease activity. Although cA6 and cA5 bind Csm6-2 with comparable affinity, only cA5 induces CARF domain closure, stabilizes the tetramer, and remodels the active site in the HEPN domain. In contrast, the sixth AMP of cA6 imposes significant steric hindrance on CARF domain movement, preventing its closure and subsequent allosteric activation. These findings expand our understanding of the cOA signaling diversity and specific cOA recognition mechanisms in type III CRISPR immunity.
- College of Chemistry, Fuzhou University, Fuzhou, China.
Organizational Affiliation: