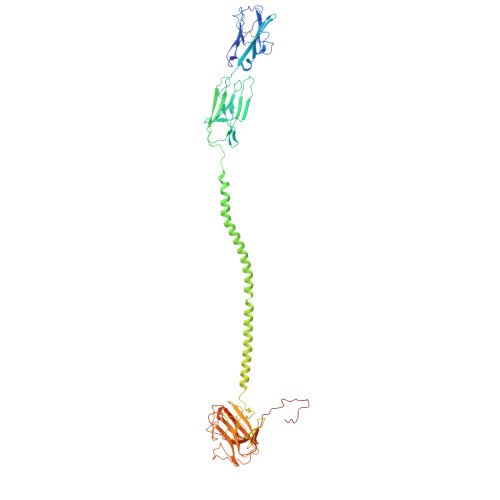
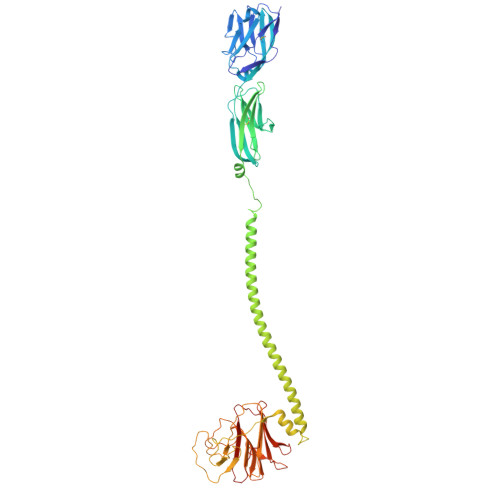
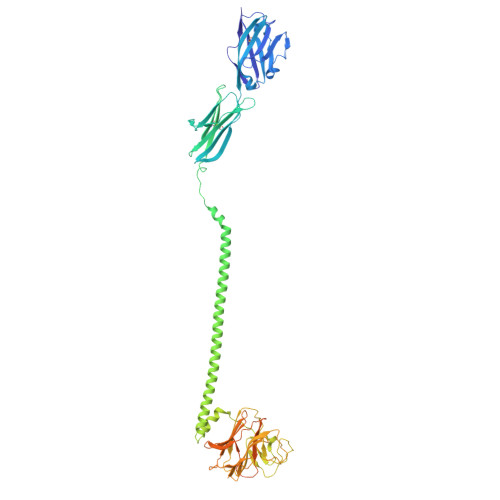
Phosphoantigen-induced inside-out stabilization of butyrophilin receptor complexes drives dimerization-dependent gamma delta TCR activation.
Zhu, Y., Gao, W., Zheng, J., Bai, Y., Tian, X., Huang, T., Lu, Z., Dong, D., Zhang, A., Guo, C., Huang, Z.(2025) Immunity 58: 1646-1659.e5
- PubMed: 40334665
- DOI: https://doi.org/10.1016/j.immuni.2025.04.012
- Primary Citation of Related Structures:
8ZA6, 8ZA9, 8ZAA, 8ZAB, 8ZD4, 8ZYR, 9II6, 9IIK, 9J5J, 9J5M - PubMed Abstract:
Phosphoantigens (pAgs), produced by infected or cancer cells, trigger the assembly of a membrane receptor complex comprising butyrophilin (BTN) members BTN3A1 and BTN2A1, leading to the activation of γδ T cells. BTN3A2 or BTN3A3 forms heteromers with BTN3A1, exhibiting higher γδ T cell receptor (TCR)-stimulating activity than BTN3A1 homomers. Cryoelectron microscopy (cryo-EM) structure reveals a pAg-induced BTN2A1-BTN3A1 heterotetramer with a 2:2 stoichiometry, stabilized by interactions between the intracellular B30.2 domains and the extracellular immunoglobulin V (IgV) domains. BTN3A2 or BTN3A3 heterodimerizes with BTN3A1, forming a pAg-induced tetrameric complex with BTN2A1. However, BTN3A1 heterodimers are more stable than BTN3A1 homodimers in this interaction. Cryo-EM reveals that BTN2A1-BTN3A1-BTN3A2 binds two γδ TCR ectodomains, with one being sandwiched between the IgV domains of BTN2A1 and BTN3A2, while the other interacts with the free BTN2A1 IgV in the complex, as evidenced by functional data. Together, our findings uncover the mechanism of ligand-induced inside-out stabilization of BTN receptor complexes for dimeric activation of γδ TCR.
- HIT Center for Life Sciences, School of Life Science and Technology, Harbin Institute of Technology, Harbin 150001, China.
Organizational Affiliation: