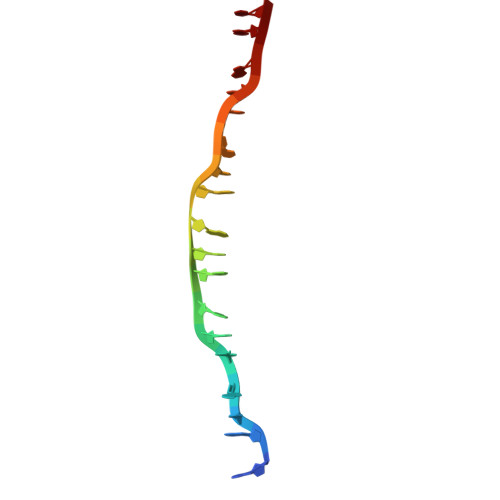
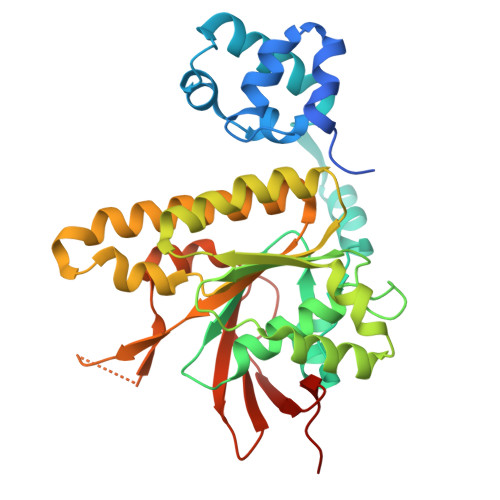
Structures and molecular mechanisms of RAD54B in modulating homologous recombination.
Liang, P., Tye, S., Ertl da Costa, J., Maharshi, N., Argunhan, B., Kuhlen, L., Battley, M., McCormack, E.A., Heyer, W.D., Lobrich, M., Zhang, X.(2026) bioRxiv
- PubMed: 41929041
- DOI: https://doi.org/10.64898/2026.03.22.713441
- Primary Citation Related Structures:
9SRZ, 9SSL, 9TRL, 9TRM, 9TYY - PubMed Abstract:
Genome stability is essential for cellular viability yet constantly threatened by endogenous and exogenous DNA-damaging agents. Among these, DNA double-strand breaks (DSBs) are particularly harmful and in S/G2 phases are faithfully repaired through homologous recombination (HR), a high-fidelity pathway utilising homologous sequences in sister chromatin. The RAD51 recombinase forms nucleoprotein filaments on single-stranded DNA (ssDNA) to mediate homology search, strand invasion and subsequent D-loop formation that leads to DNA synthesis and repair. The efficiency of HR depends on precise regulation of RAD51 filament dynamics by accessory factors, including RAD54 and RAD54B, which belong to the SWI2/SNF2-family DNA translocases. While RAD54 is well-characterized, RAD54B's molecular functions remain poorly understood. Here, we define RAD54B's role in HR using cryo-electron microscopy, mutagenesis, biochemical and cellular assays. We show that RAD54B stabilizes RAD51-DNA filaments, inhibits RAD51 ATPase activity, and promotes strand invasion, D-loop formation and strand exchange. The N-terminal domain (NTD) alone supports filament stabilization and strand exchange, while the C-terminal ATPase domain is required for D-loop formation. Structural and biochemical analyses reveal three RAD51-interacting sites within the NTD and a unique domain (β-domain) that bridges RAD51 protomers and contacts donor dsDNA. This β-domain also regulates RAD54B's ATPase activity and higher-order oligomer organization on dsDNA. Cellular assays reveal that the NTD RAD51-interacting sites as well as the β-domain are required for repairing camptothecin-induced DSBs by HR in human cells. Our findings uncover a modular architecture and mechanistic framework for RAD54B function in HR, highlighting its critical role in genome maintenance. cryoEM structure of RAD54B in complex with RAD51-DNA complexRAD54B uses three sites to interact with RAD51, including a previously unrecognised β-domain that bridges distal RAD51 protomers.The β-domain plays multiple crucial roles including regulating filament stability, RAD54B ATPase activity and RAD54B higher order assembly on DNA.RAD54B employs a modular mechanism, with the N-terminal region engaing and stabilising RAD51 filaments, capturing of the homologous strands, whereas the ATPase motor domainrequired for homology search and strand invasion.RAD54B N-terminus and β-domain are essential for HR-mediated repair of camptothecin-induced breaks in human cells.